Sex Testing is about accuracy. Failure is not an option. For this reason one needs to thoroughly understand what a given assay actually targets in the cannabis genome and the validation set behind a given product or service offering. To accomplish this we have Whole genome sequenced 12 Female Cannabis Strains and 2 Male cannabis strains.
By mapping all of the male sequences against female sequences we in silico distill a list of sequences in the male plants that do not exist in the 12 female genomes. We then assemble these male specific reads into a Y Chromosome.
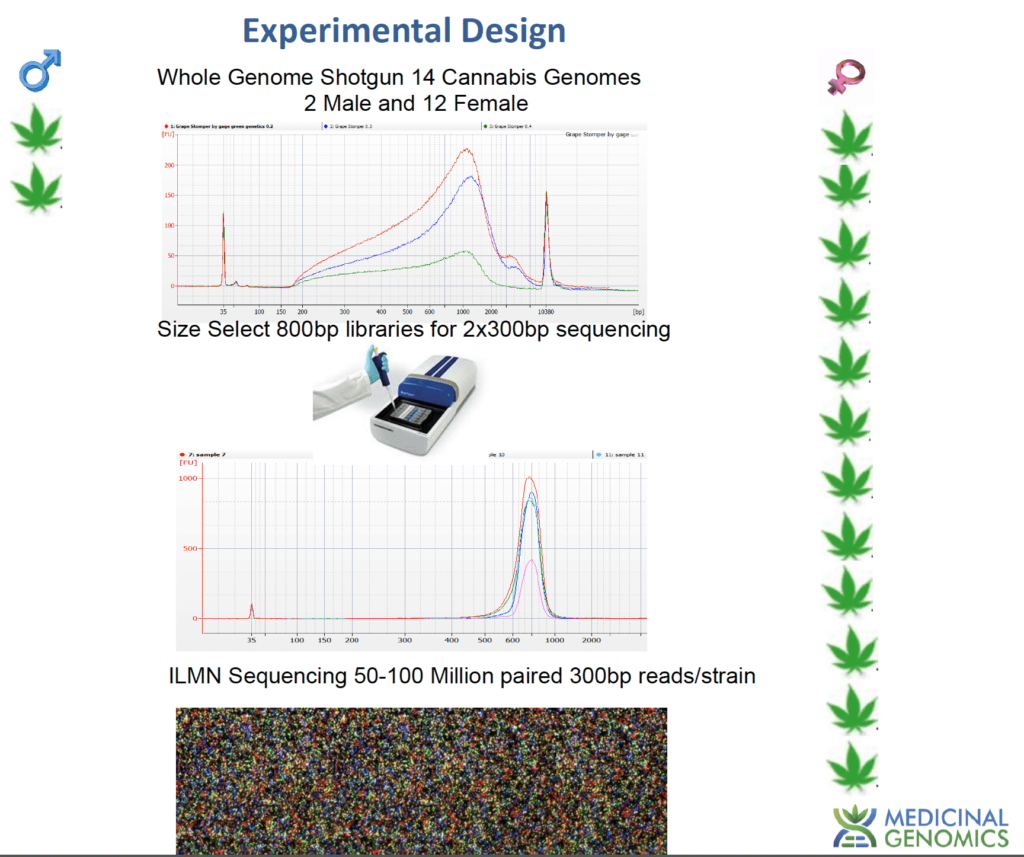
Assembled contigs are then filtered for bacteria, mold and known cannabis endophytes. These sequences are further filtered for heterozygosity to deliver hundreds of target qPCR loci. These loci were screened for the best performance in dozens of unrelated males.
To further confirm the conservation of our loci, we sequenced a cannabis cultivar that is thought to be the most genetically distant cannabis cultivar known to man (In Press). This cultivar revealed 11% of the Y chromosome that is absent in distant lineages and underscores the importance of thorough validation and screening of cannabis genetics before one believes a male specific assay will be fail safe in practical use. qPCR assays can then target the Y chromosome and an autosomal cannabinoid synthase gene in a multiplexed assay to provide male vs female answers in hours from a leaf punch.
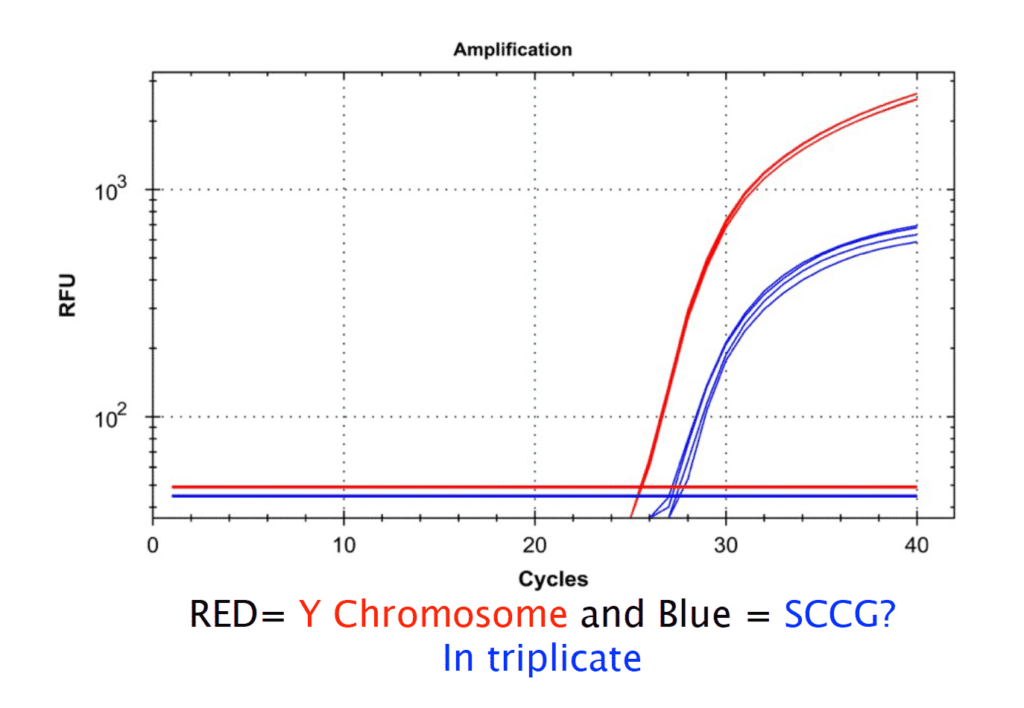
For more about the Male Cannabis Genome Project see THIS LINK.
FAQ-
Why are you not using the published Mandolino Primer sets?
In addition to Mandolino et al. publishing pioneering work, they also brilliantly figured out part of this problem before we had access to whole genome sequencing of many cannabis cultivars. The challenge in doing this without nextgen sequencing cannot be underscored enough and like all inventions the next generation has the benefit of standing on the shoulders of giants. Further sequencing of the Mandolino published MADC2 regions have shown these regions to be problematic targets for qPCR. The main problem with the Mandolino primer sets is that they target a repeat in the genome that is not unique to Males. A single base variants in this region provides one extra band for most males. Most males is the key issues here. When failure is not an option repeats with 500 copies in the female genome cannot be part of a qPCR product offering to identify males. Too little is know about these mobile elements to bet on them for large crops at risk of pollination. Ask your test provider what they target and how they validated their male test before they blame you for “hermi”ing your plants.
Below is depiction of the MADC2 region after mapping reads to the loci. This was a 30X male genome that ended up with 17,000X coverage over the MADC2 loci and many polymorphisms (vertical lines) with very few paired reads agreeing on the full length nature of the repeat. This explains the complicated banding patterns described in the Mandolino et al paper. The blue brackets below represent the primer locations. The polymorphisms under the primers are a concern for qPCR assays.
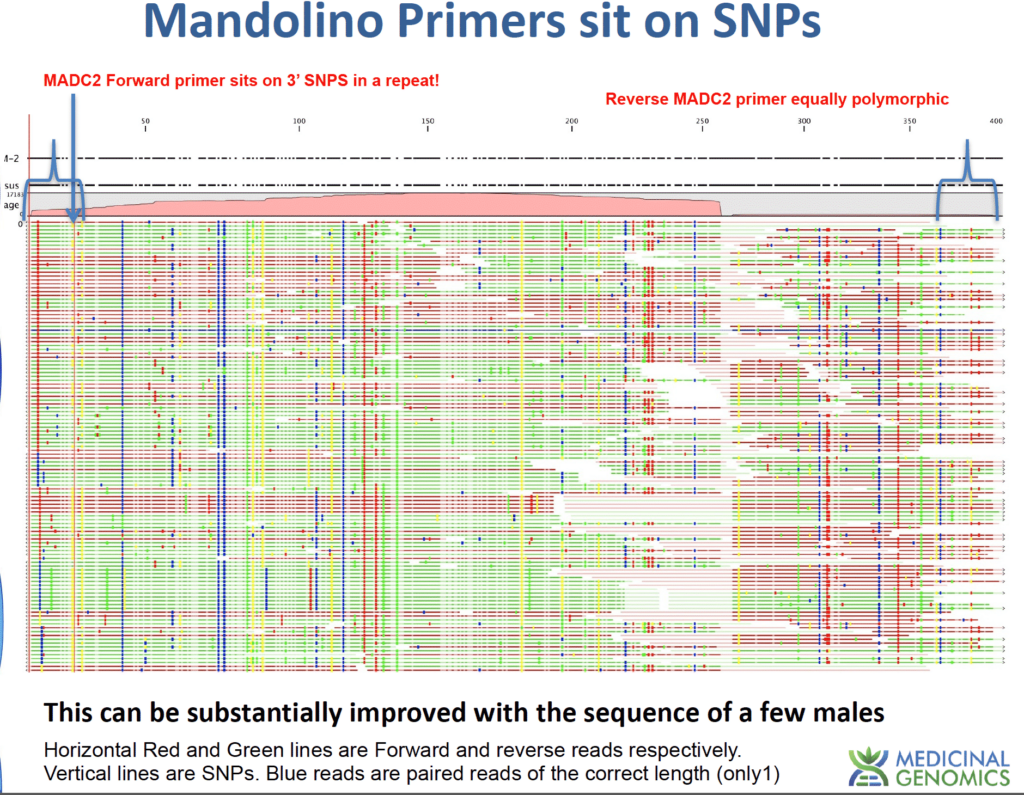
To finally confirm our Male markers we sequenced a male genome from the most distant Cannabis species we could find. 11% of the Y chromosome we had under target for the Male project did not exist in this distant species. Redesigning our assay to adequately call this Feral hemp strain has provided 100% accuracy calling male plants to date. We encourage folks to send us SenSATIVAx DNA from males with photographic confirmation of pollen sacks andwe will increase the N number.
Another set of samples from Salmon Creek in Southern Humboldt County tested perfectly in June of 2015. Included in this study is a set of unknowns yet to express Sex.
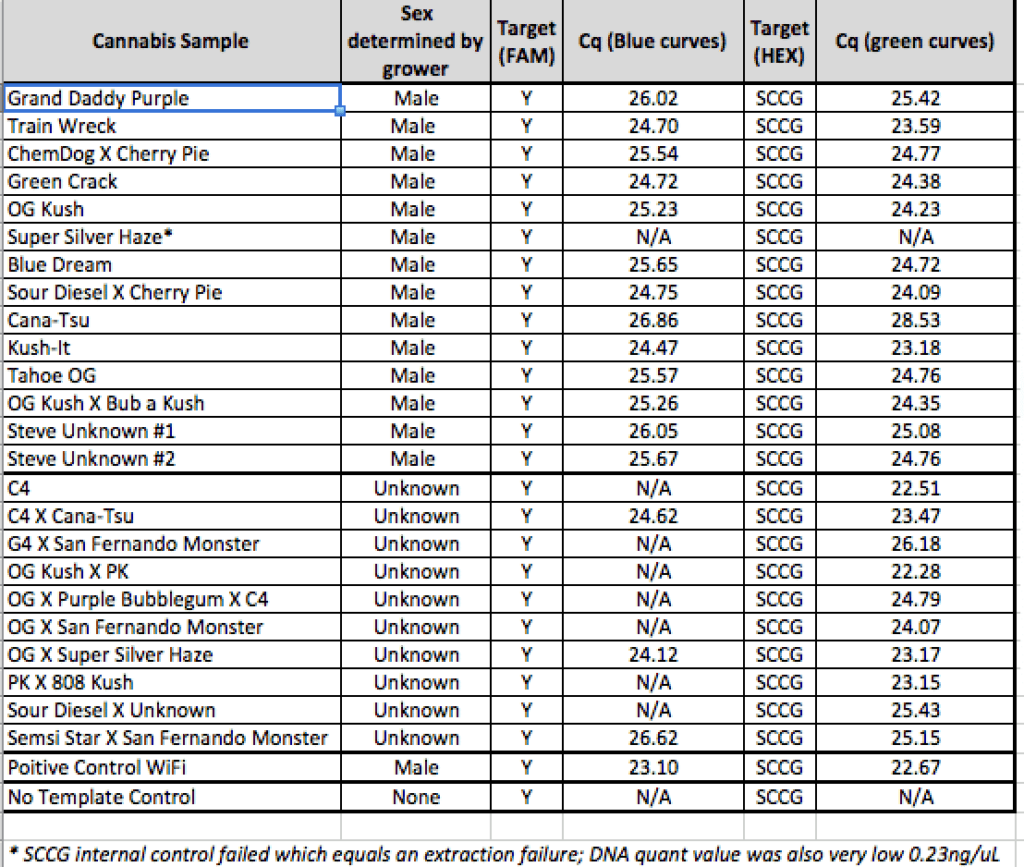
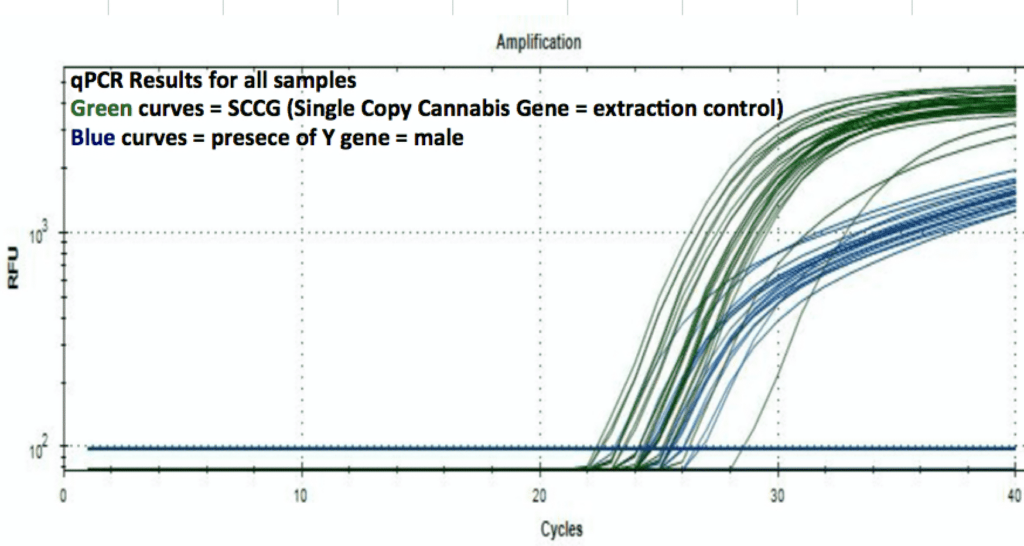
Beta Site Validation Data on Roche 480 cycler
100% accuracy
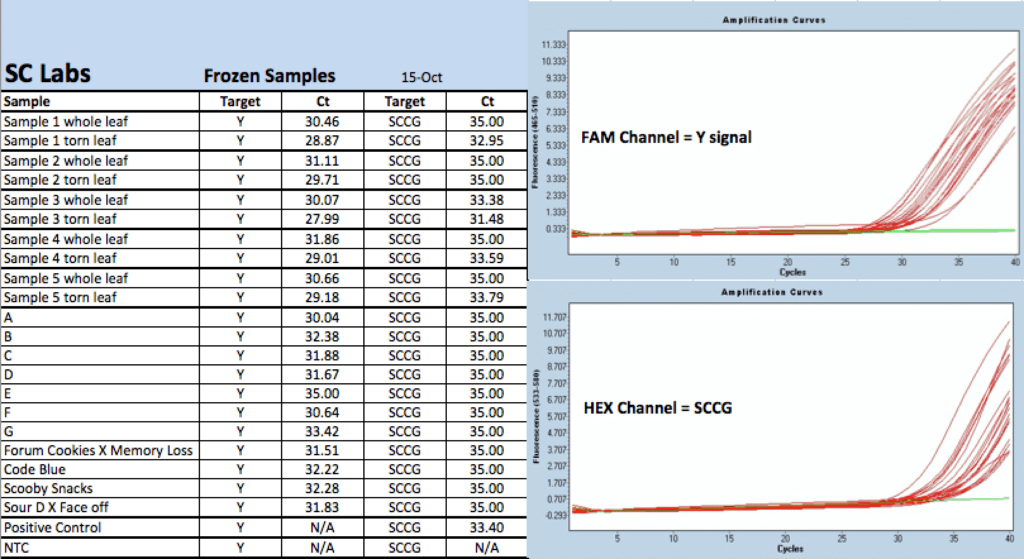
Expanded Trial
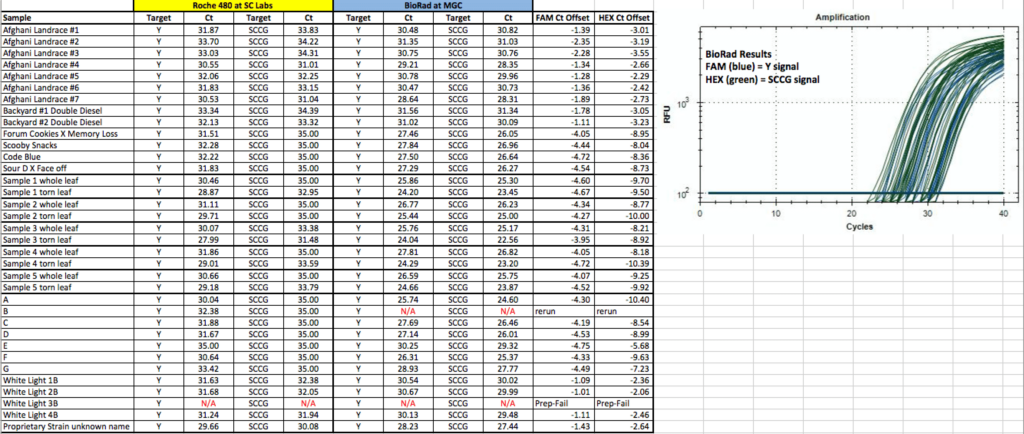
Suspected Prep Failures Re-Preppped and ReRun successfully on Bio-Rad.

The SCCG signals are a bit late. More work on the Roche 480 maybe required to optimize the HEX channel as Roche disclaimers signals greater than CT of 35.
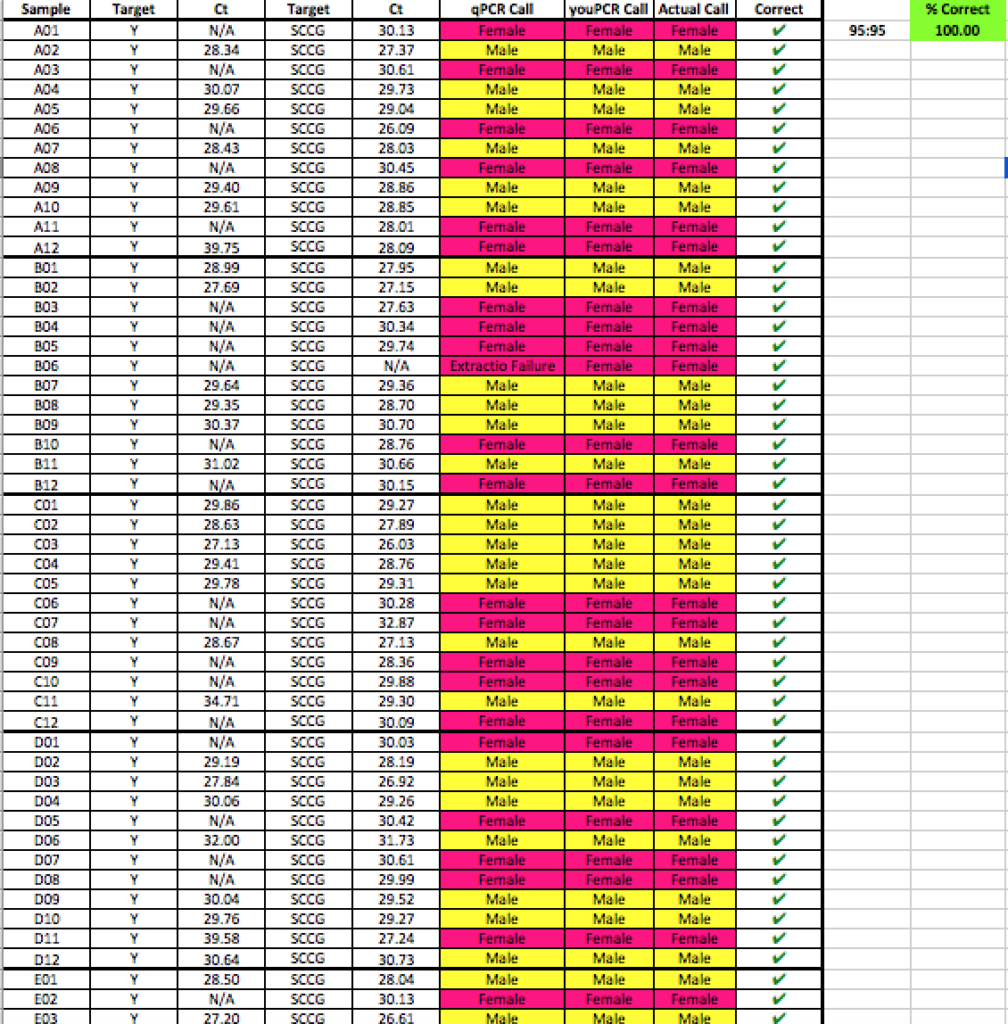
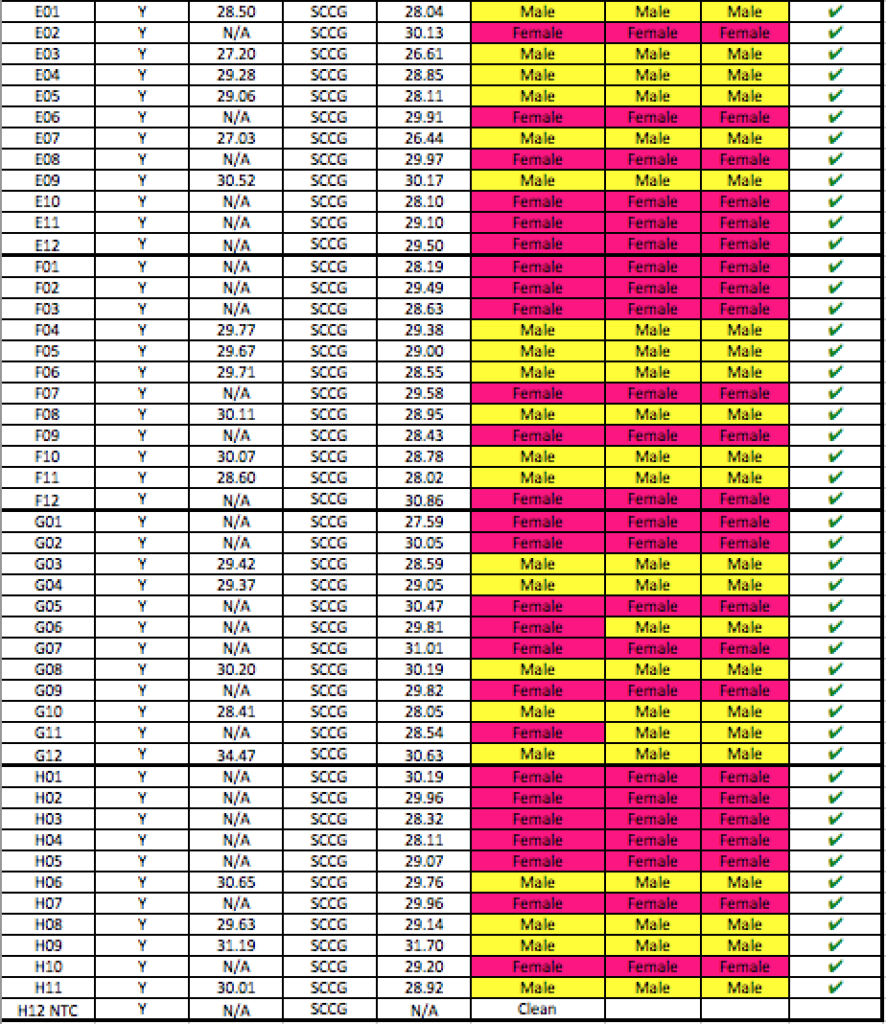
Below is a blinded Validation performed in a seed to sale system at The Slater Center in Providence RI.
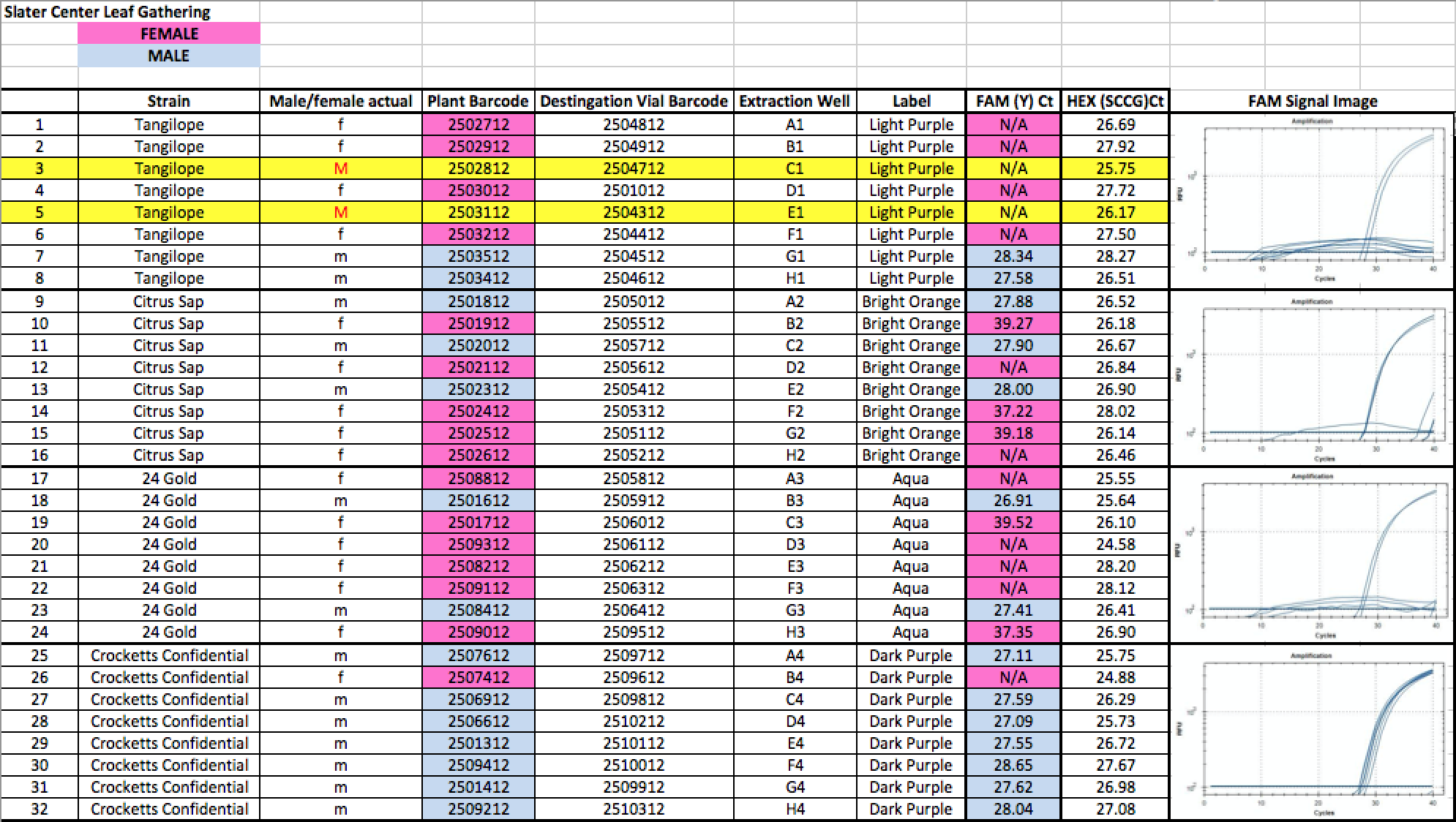
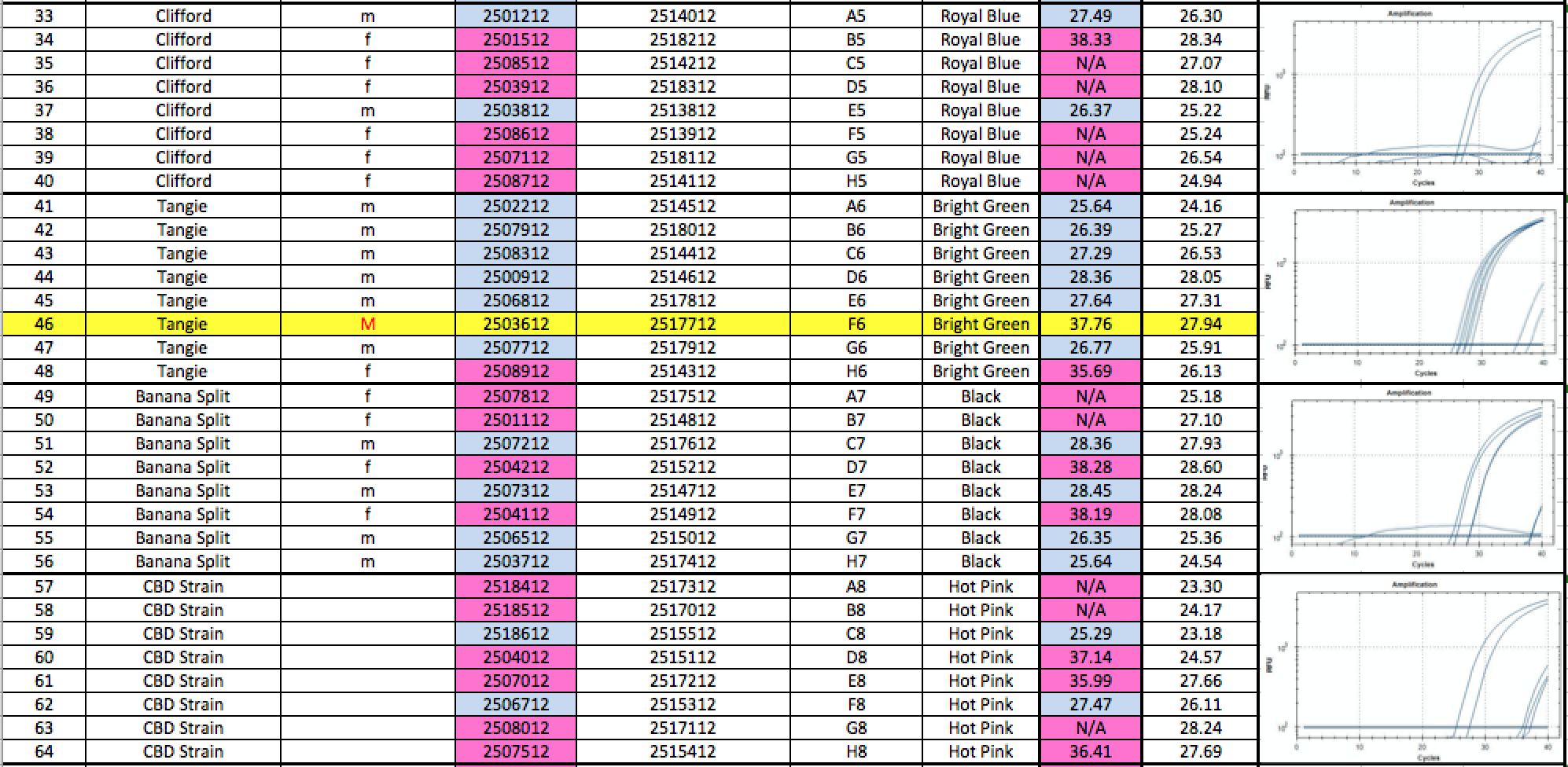
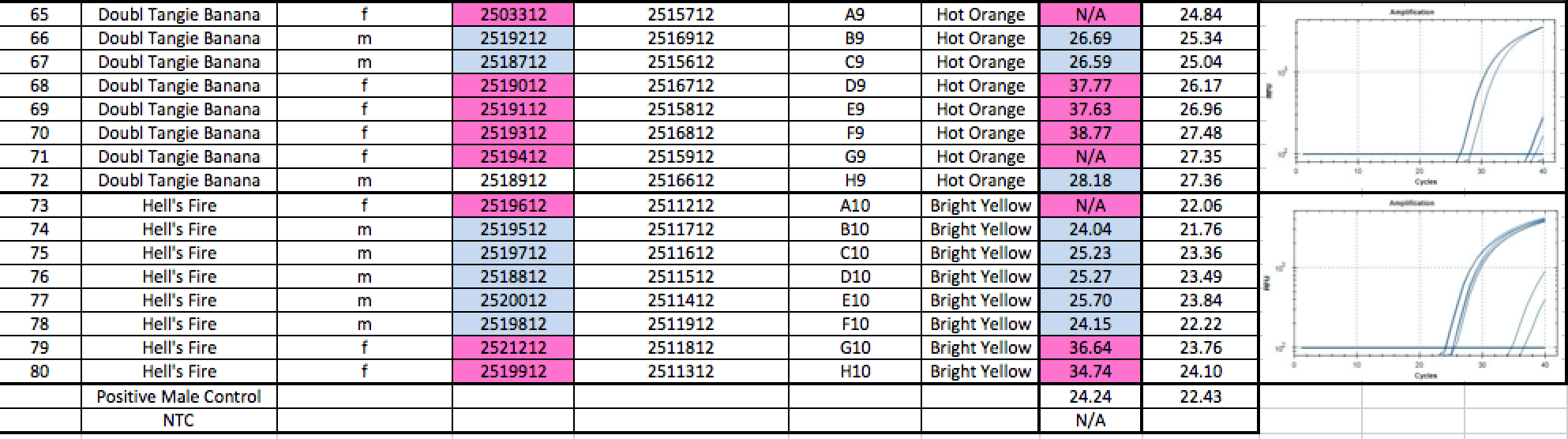
A few siblings in the Tangie and Tangilope lines are being sequenced to explore the possibility of SNPs under our primers for these lines. Its interesting that most of the other male siblings qPCR’d successfully as Males.
Any validation should include samples from geographically distinct regions to ensure complexity in the sampling. The value of the internal controls is evident. When samples fail to amplify a gene in the cannabinoid synthase pathway, the DNA prep is suspect and the lack of Y signal cannot be inferred as Female.
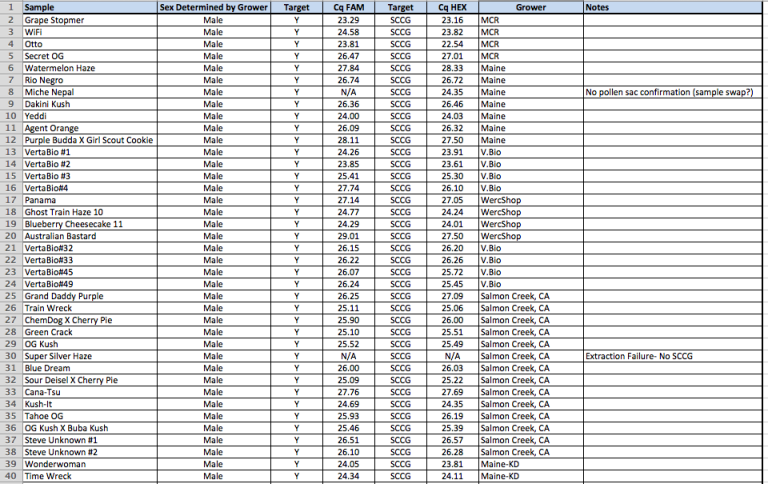
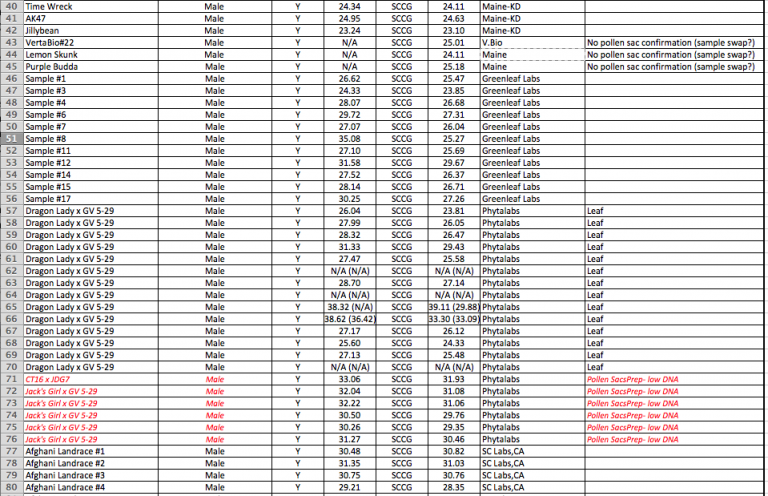
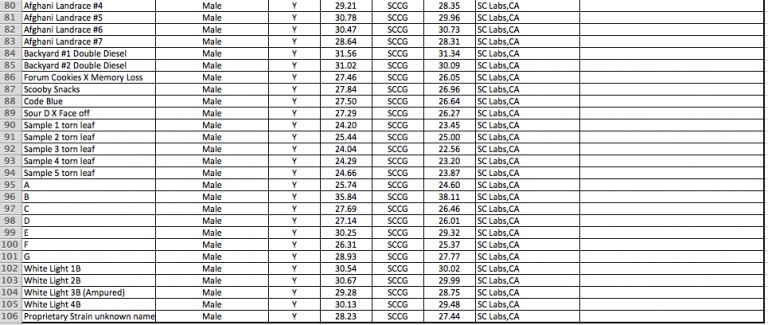
YouPCR Sex Test Validation with Colorado Seeds
Batch #2
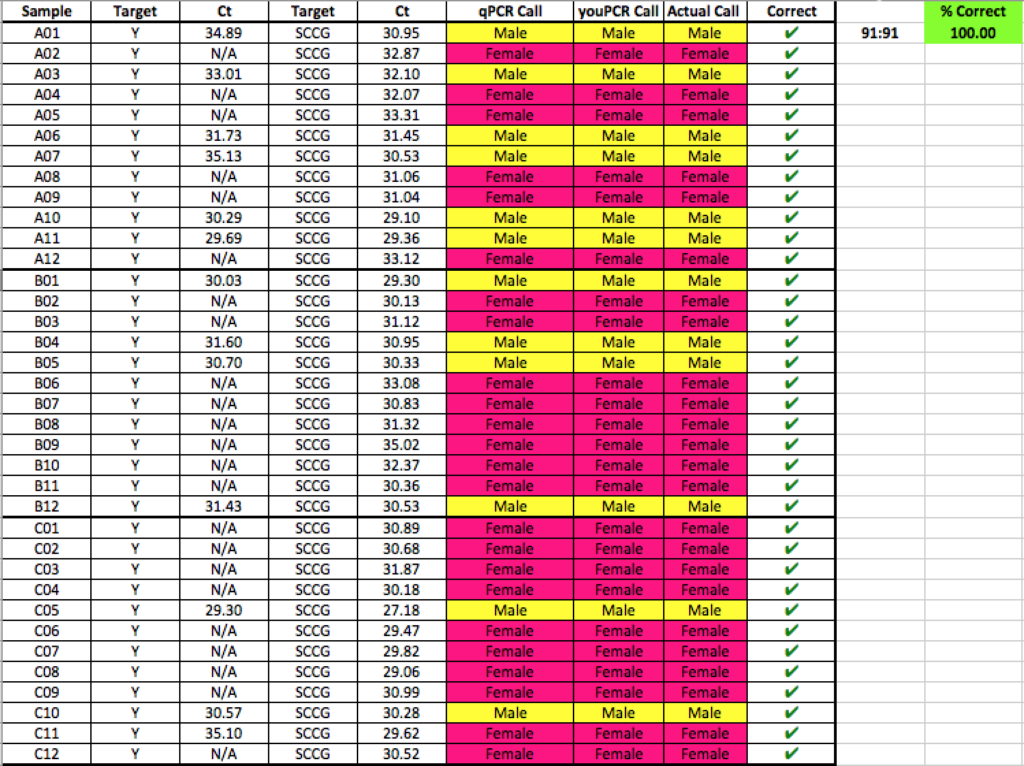
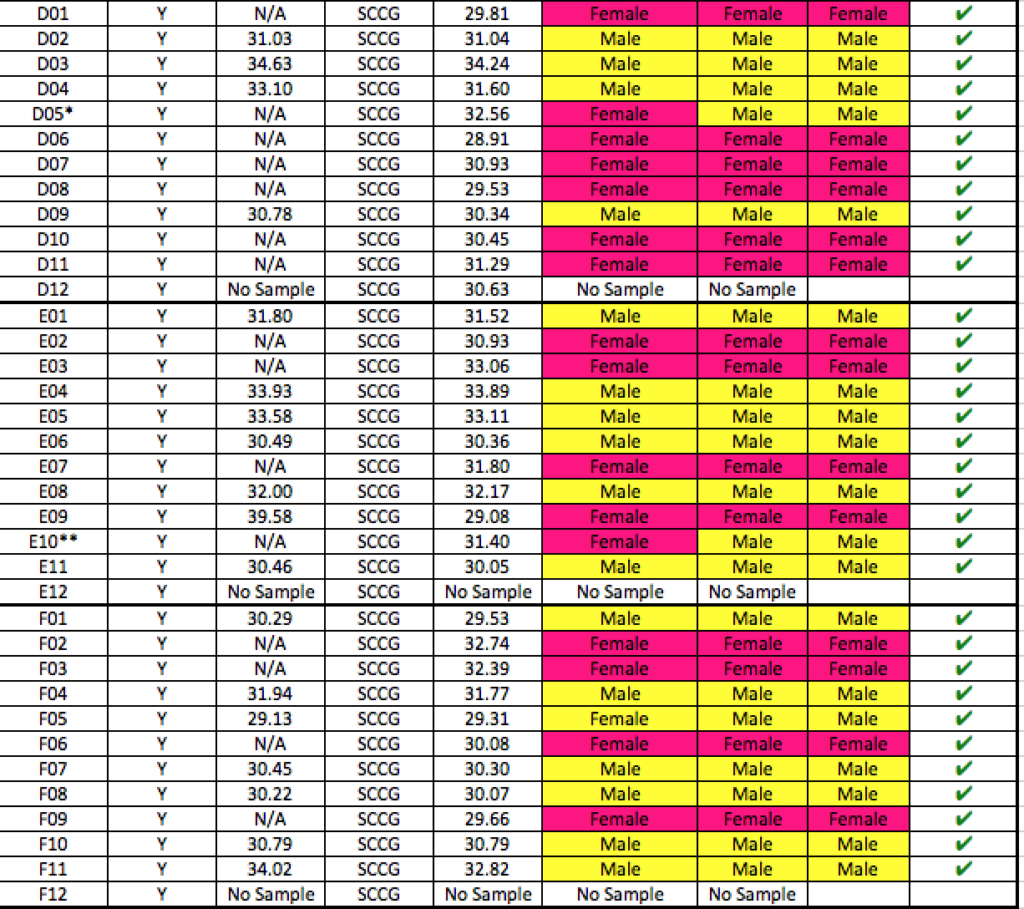
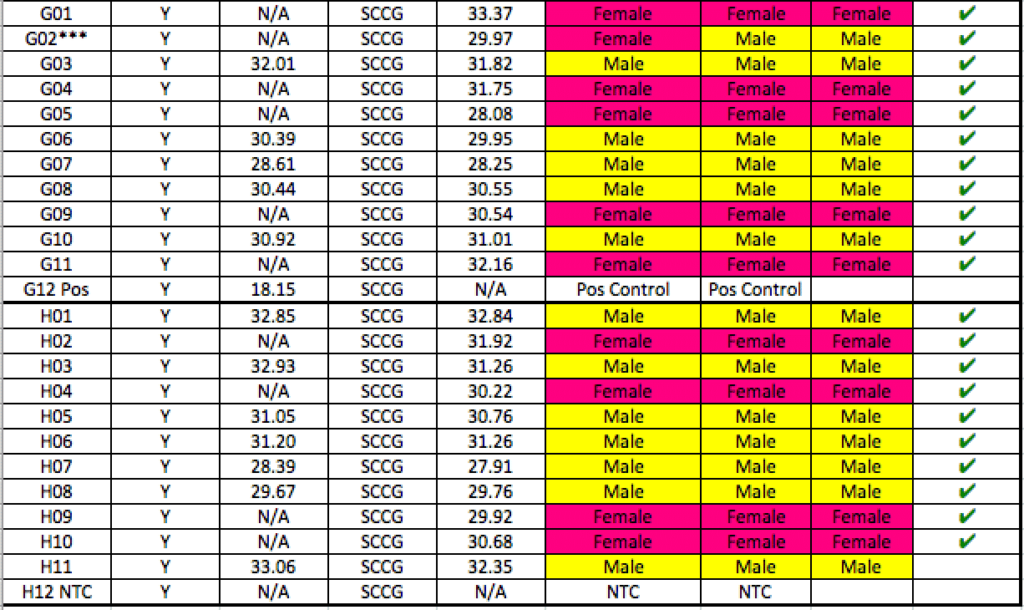
Batch #1
©2024 MEDICINAL GENOMICS CORPS
Copyright all right reserved.
Signup to receive email updates on blog posts, podcasts, webinars, new products, and more!
Join us at the CannMed 24 Innovation and Investment Summit, an exclusive experience of discovery, development, and networking with innovators from around the world.